Trans
What is Translational Science?
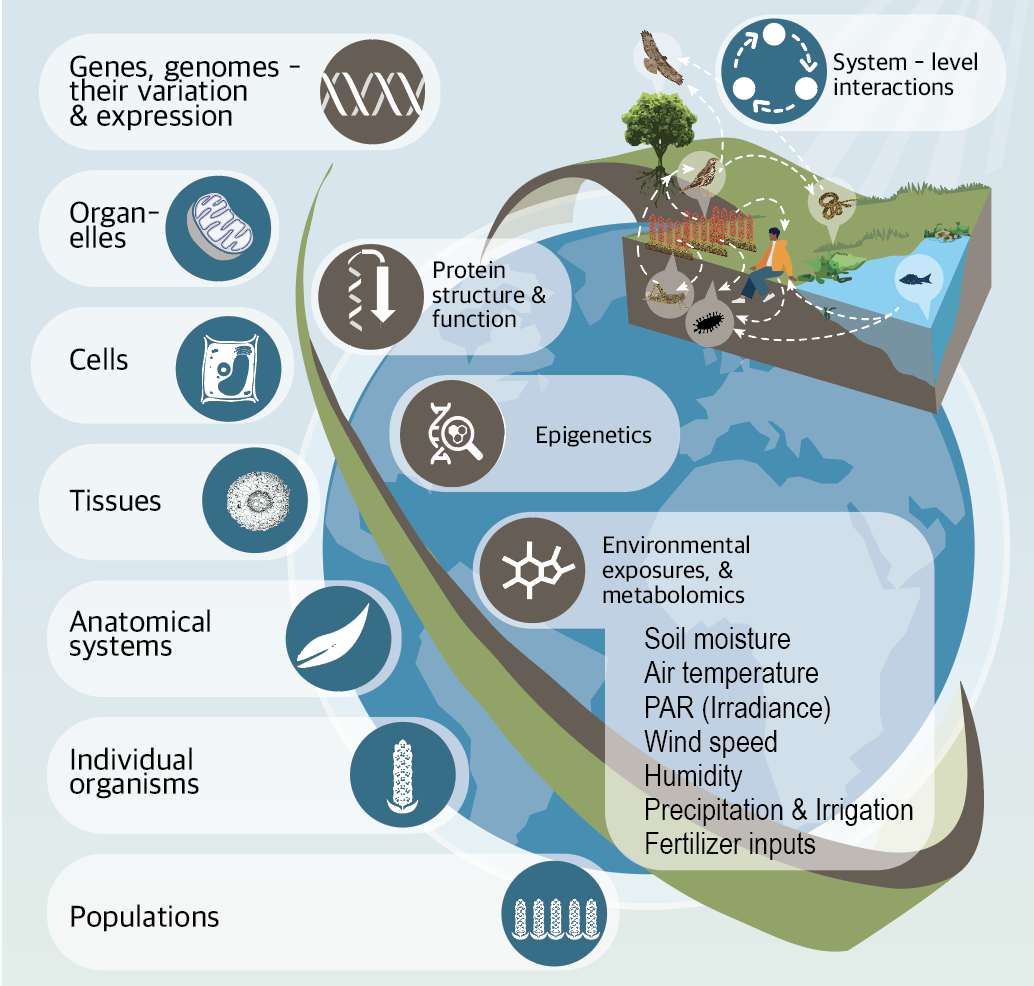
Translational science breaks down traditional silos between discreet disciplines of science empowering innovation. At the forefront of this emerging field, the Translational and Integrative Sciences Lab (TISLab) brings together domain experts to study phenotype data—information about observable traits—to unlock new insights. TISLab’s work develops new understanding and amplifies discovery in areas such as rare disease diagnosis in both plants and humans, accelerates cancer research, and deepens our understanding of how exposures and other factors influence health. Because of its inherently cross-disciplinary nature, TISLab’s research thrives on collaboration and partnership.
